---
title: "One Intervention, System-Wide Impact"
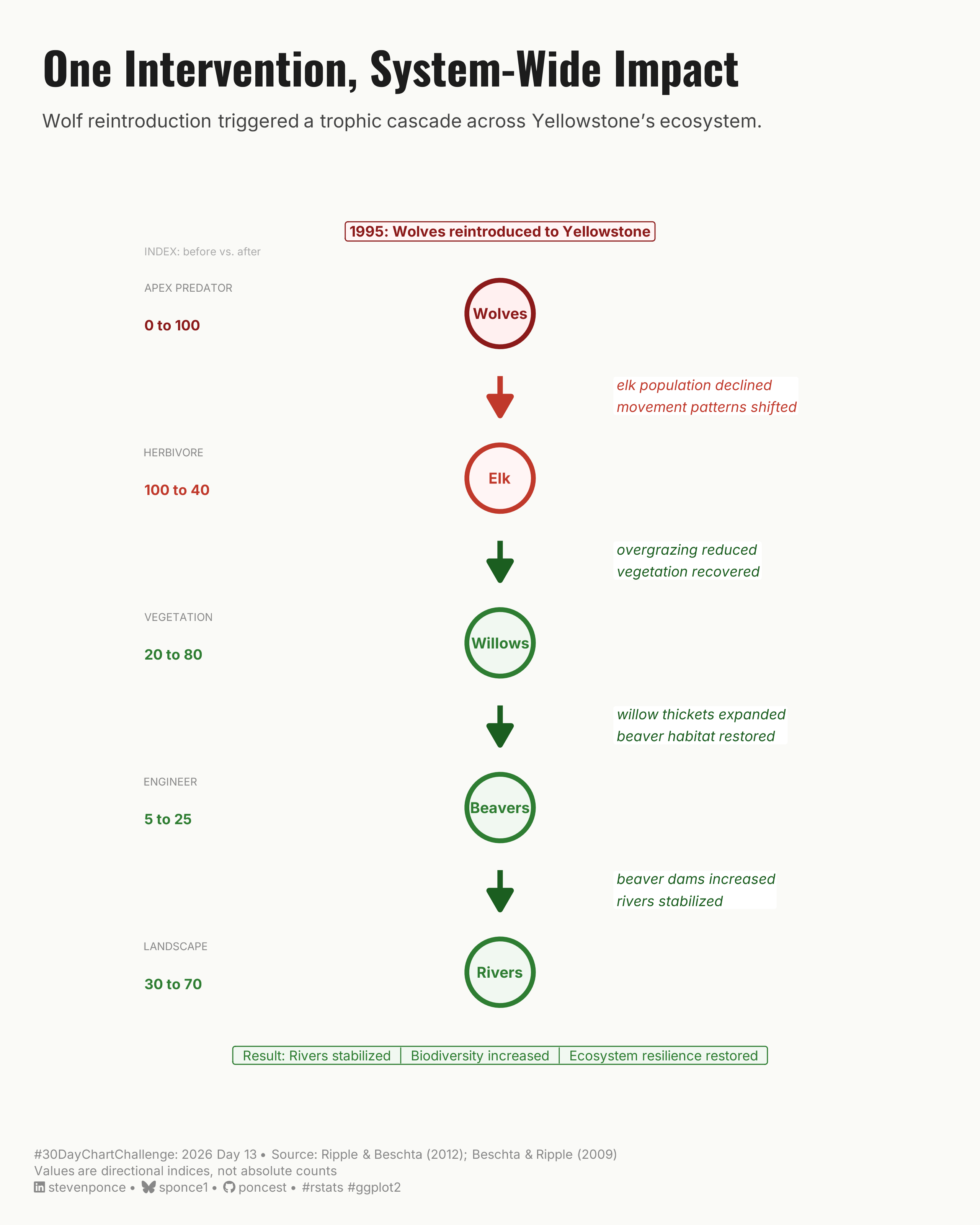
subtitle: "Wolf reintroduction triggered a trophic cascade across Yellowstone's ecosystem."
description: "A vertical trophic cascade diagram showing how wolf reintroduction in Yellowstone in 1995 triggered system-wide ecological recovery across five interdependent levels — from apex predator to river systems. Built with ggplot2 using curated directional indices based on Ripple & Beschta (2012)."
date: "2026-04-13"
author:
- name: "Steven Ponce"
url: "https://stevenponce.netlify.app"
citation:
url: "https://stevenponce.netlify.app/data_visualizations/30DayChartChallenge/2026/30dcc_2026_13.html"
categories: ["30DayChartChallenge", "Data Visualization", "R Programming", "2026"]
tags: [
"30DayChartChallenge",
"Relationships",
"Ecosystems",
"Trophic Cascade",
"Node-Arrow Diagram",
"Yellowstone",
"Wolf Reintroduction",
"Ecology",
"Causal Diagram",
"System Diagram",
"ggplot2",
"Annotated Chart",
"Conservation"
]
image: "thumbnails/30dcc_2026_13.png"
format:
html:
toc: true
toc-depth: 5
code-link: true
code-fold: true
code-tools: true
code-summary: "Show code"
self-contained: true
theme:
light: [flatly, assets/styling/custom_styles.scss]
dark: [darkly, assets/styling/custom_styles_dark.scss]
editor_options:
chunk_output_type: inline
execute:
freeze: true
cache: true
error: false
message: false
warning: false
eval: true
---
{#fig-1}
### [**Steps to Create this Graphic**]{.mark}
#### [1. Load Packages & Setup]{.smallcaps}
```{r}
#| label: load
#| warning: false
#| message: false
#| results: "hide"
## 1. LOAD PACKAGES & SETUP ----
suppressPackageStartupMessages({
pacman::p_load(
tidyverse, ggtext, showtext,
janitor, scales, glue
)
})
### |- figure size ----
camcorder::gg_record(
dir = here::here("temp_plots"),
device = "png",
width = 10,
height = 8,
units = "in",
dpi = 320
)
# Source utility functions
suppressMessages(source(here::here("R/utils/fonts.R")))
source(here::here("R/utils/social_icons.R"))
source(here::here("R/utils/image_utils.R"))
source(here::here("R/themes/base_theme.R"))
```
#### [2. Read in the Data]{.smallcaps}
```{r}
#| label: read
#| include: true
#| eval: true
#| warning: false
### |- trophic nodes ----
# Each entity = one level in the Yellowstone cascade
# y positions are fixed for vertical layout (top → bottom)
nodes <- tribble(
~entity, ~trophic_level, ~y, ~before, ~after, ~direction,
"Wolves", "Apex Predator", 4.0, 0, 100, "restored",
"Elk", "Herbivore", 3.0, 100, 40, "reduced",
"Willows", "Vegetation", 2.0, 20, 80, "recovered",
"Beavers", "Engineer", 1.0, 5, 25, "recovered",
"Rivers", "Landscape", 0.0, 30, 70, "stabilized"
)
### |- arrows (edges) ----,
# Each row = one causal link between consecutive trophic levels
# label = annotation text on the arrow
# effect = positive or negative pressure
edges <- tribble(
~from, ~to, ~y_from, ~y_to, ~effect, ~label,
"Wolves", "Elk", 5.0, 3.8, "negative", "elk population declined\nmovement patterns shifted",
"Elk", "Willows", 3.8, 2.6, "positive", "overgrazing reduced\nvegetation recovered",
"Willows", "Beavers", 2.6, 1.4, "positive", "willow thickets expanded\nbeaver habitat restored",
"Beavers", "Rivers", 1.4, 0.2, "positive", "beaver dams increased\nrivers stabilized"
)
```
#### [3. Examine the Data]{.smallcaps}
```{r}
#| label: examine
#| include: true
#| eval: true
#| results: 'hide'
#| warning: false
glimpse(nodes)
```
#### [4. Tidy Data]{.smallcaps}
```{r}
#| label: tidy
#| warning: false
### |- x positions ----
x_node <- 0.0
x_seg <- 0.0
x_label_right <- 0.28
x_left <- -0.88
node_r <- 0.38
```
#### [5. Visualization Parameters]{.smallcaps}
```{r}
#| label: params
#| include: true
#| warning: false
### |- plot aesthetics ----
colors <- get_theme_colors(
palette = list(
col_wolf = "#8B1A1A",
col_elk = "#C0392B",
col_recovered = "#2E7D32",
col_neutral = "#455A64",
col_arrow_neg = "#C0392B",
col_arrow_pos = "#1B5E20",
col_label_bg = "#FFFFFF",
bg_color = "#FAFAF7",
text_color = "#1C1C1C",
grid_color = "#E8E8E4"
)
)
### |- node color assignment ----
nodes <- nodes |>
mutate(
node_color = case_when(
entity == "Wolves" ~ colors$palette$col_wolf,
entity == "Elk" ~ colors$palette$col_elk,
TRUE ~ colors$palette$col_recovered
),
node_color_text = case_when(
entity == "Wolves" ~ colors$palette$col_wolf,
entity == "Elk" ~ colors$palette$col_elk,
TRUE ~ colors$palette$col_recovered
),
node_fill = case_when(
entity == "Wolves" ~ "#FFF0F0",
entity == "Elk" ~ "#FFF5F5",
TRUE ~ "#F1F8F1"
)
)
edges <- edges |>
mutate(
y_from = case_when(
from == "Wolves" ~ 4.0,
from == "Elk" ~ 3.0,
from == "Willows" ~ 2.0,
from == "Beavers" ~ 1.0
),
y_to = case_when(
to == "Elk" ~ 3.0,
to == "Willows" ~ 2.0,
to == "Beavers" ~ 1.0,
to == "Rivers" ~ 0.0
),
arrow_color = if_else(
effect == "negative",
colors$palette$col_arrow_neg,
colors$palette$col_arrow_pos)
)
### |- titles and caption ----
title_text <- "One Intervention, System-Wide Impact"
subtitle_text <- "Wolf reintroduction triggered a trophic cascade across Yellowstone's ecosystem."
caption_text <- create_dcc_caption(
dcc_year = 2026,
dcc_day = 13,
source_text = "Ripple & Beschta (2012); Beschta & Ripple (2009)<br>Values are directional indices, not absolute counts"
)
### |- fonts ----
setup_fonts()
fonts <- get_font_families()
```
#### [6. Plot]{.smallcaps}
```{r}
#| label: plot
#| warning: false
### |- main plot ----
p <- ggplot() +
# trophic level labels
geom_text(
data = nodes,
aes(x = x_left, y = y + 0.18, label = str_to_upper(trophic_level)),
hjust = 0,
vjust = 1,
size = 2.2,
color = "#888888",
family = fonts$text,
fontface = "plain"
) +
# before/after index
geom_text(
data = nodes,
aes(
x = x_left, y = y - 0.04,
label = glue("{before} to {after}"),
color = I(node_color)
),
hjust = 0,
vjust = 1,
size = 2.9,
family = fonts$text,
fontface = "bold"
) +
# straight arrow segments
geom_segment(
data = edges,
aes(
y = y_from - node_r,
yend = y_to + node_r,
color = I(arrow_color)
),
x = 0,
xend = 0,
linewidth = 1.5,
arrow = arrow(length = unit(0.18, "inches"), type = "closed", ends = "last"),
lineend = "butt"
) +
# edge annotation labels
geom_label(
data = edges,
aes(
x = x_label_right,
y = (y_from + y_to) / 2,
label = label,
color = I(arrow_color)
),
hjust = 0,
vjust = 0.5,
size = 2.9,
family = fonts$text,
fontface = "italic",
fill = colors$palette$col_label_bg,
linewidth = 0,
label.padding = unit(0.18, "lines"),
lineheight = 1.3
) +
# node circles
geom_point(
data = nodes,
aes(y = y, color = I(node_color), fill = I(node_fill)),
x = 0,
size = 17,
shape = 21,
stroke = 2.0
) +
# node entity labels
geom_text(
data = nodes,
aes(y = y, label = entity, color = I(node_color)),
x = 0,
size = 3.1,
family = fonts$text,
fontface = "bold"
) +
# "INDEX" column header
annotate(
"text",
x = x_left, y = 4.40,
label = "INDEX: before vs. after",
hjust = 0,
vjust = 1,
size = 2.2,
color = "#AAAAAA",
family = fonts$text
) +
# intervention callout
annotate(
"label",
x = 0, y = 4.44,
label = "1995: Wolves reintroduced to Yellowstone",
hjust = 0.5,
vjust = 0,
size = 3.0,
color = colors$palette$col_wolf,
fill = "#FFF5F5",
linewidth = 0.3,
family = fonts$text,
fontface = "bold"
) +
# outcome callout
annotate(
"label",
x = 0, y = -0.45,
label = "Result: Rivers stabilized | Biodiversity increased | Ecosystem resilience restored",
hjust = 0.5,
vjust = 1,
size = 2.8,
color = colors$palette$col_recovered,
fill = "#F1F8F1",
linewidth = 0.3,
family = fonts$text
) +
# scales
scale_x_continuous(limits = c(-1.05, 1)) +
scale_y_continuous(limits = c(-0.72, 4.75)) +
# labs
labs(
title = title_text,
subtitle = subtitle_text,
caption = caption_text
) +
# theme
theme_void() +
theme(
plot.background = element_rect(fill = colors$palette$bg_color, color = NA),
panel.background = element_rect(fill = colors$palette$bg_color, color = NA),
plot.title = element_text(
family = fonts$title, face = "bold",
size = 26, color = colors$palette$text_color,
margin = margin(t = 10, b = 6, l = 5)
),
plot.subtitle = element_textbox_simple(
family = fonts$text,
size = 11,
color = "#444444",
lineheight = 1.3,
margin = margin(t = 5, b = 8, l = 5)
),
plot.caption = element_textbox_simple(
family = fonts$text,
size = 7.5,
color = "#888888",
lineheight = 1.3,
margin = margin(t = 8, b = 6)
),
plot.margin = margin(t = 20, r = 20, b = 12, l = 20)
)
```
#### [7. Save]{.smallcaps}
```{r}
#| label: save
#| warning: false
### |- plot image ----
save_plot(
p,
type = "30daychartchallenge",
year = 2026,
day = 13,
width = 8,
height = 10
)
```
#### [8. Session Info]{.smallcaps}
::: {.callout-tip collapse="true"}
##### Expand for Session Info
```{r, echo = FALSE}
#| eval: true
#| warning: false
sessionInfo()
```
:::
#### [9. GitHub Repository]{.smallcaps}
::: {.callout-tip collapse="true"}
##### Expand for GitHub Repo
The complete code for this analysis is available in [`30dcc_2026_13.qmd`](https://github.com/poncest/personal-website/blob/master/data_visualizations/TidyTuesday/2026/30dcc_2026_13.qmd).
For the full repository, [click here](https://github.com/poncest/personal-website/).
:::
#### [10. References]{.smallcaps}
::: {.callout-tip collapse="true"}
##### Expand for References
1. Data Sources:
- Ripple, W. J., & Beschta, R. L. (2012). Trophic cascades in Yellowstone: The first 15 years after wolf reintroduction. *Biological Conservation*, 145(1), 205–213. https://doi.org/10.1016/j.biocon.2011.11.005
- Beschta, R. L., & Ripple, W. J. (2009). Large predators and trophic cascades in terrestrial ecosystems of the western United States. *Biological Conservation*, 142(11), 2401–2414. https://doi.org/10.1016/j.biocon.2009.06.015
- Hebblewhite, M., White, C. A., Nietvelt, C. G., McKenzie, J. A., Hurd, T. E., Fryxell, J. M., Bayley, S. E., & Paquet, P. C. (2005). Human activity mediates a trophic cascade caused by wolves. *Ecology*, 86(8), 2135–2144. https://doi.org/10.1890/04-1269
2. Note: Index values (0–100) are directional and illustrative, constructed to reflect relative magnitude of change as reported in the literature above. They are not absolute population counts or official measurements.
:::
#### [11. Custom Functions Documentation]{.smallcaps}
::: {.callout-note collapse="true"}
##### 📦 Custom Helper Functions
This analysis uses custom functions from my personal module library for efficiency and consistency across projects.
**Functions Used:**
- **`fonts.R`**: `setup_fonts()`, `get_font_families()` - Font management with showtext
- **`social_icons.R`**: `create_social_caption()` - Generates formatted social media captions
- **`image_utils.R`**: `save_plot()` - Consistent plot saving with naming conventions
- **`base_theme.R`**: `create_base_theme()`, `extend_weekly_theme()`, `get_theme_colors()` - Custom ggplot2 themes
**Why custom functions?**\
These utilities standardize theming, fonts, and output across all my data visualizations. The core analysis (data tidying and visualization logic) uses only standard tidyverse packages.
**Source Code:**\
View all custom functions → [GitHub: R/utils](https://github.com/poncest/personal-website/tree/master/R)
:::
